bmotif: a package for counting motifs in biparite networks
Bipartite networks are widely-used to represent a diverse range of species interactions, such as pollination, herbivory, parasitism, and seed dispersal. The structure of these networks is usually characterized by calculating one or more metrics that capture different aspects of network architecture.
While these metrics capture useful properties of networks, they only consider structure at the scale of the whole network (the macro-scale) or individual species (the micro-scale).
‘Meso-scale’ structure between these scales is usually ignored, despite representing ecologically-important indirect interactions. Network motifs are a framework for capturing this meso-scale structure, and are gaining in popularity.
However, to date there is no software available in R, the most popular programming language among ecologists, for conducting motif analyses in bipartite networks.
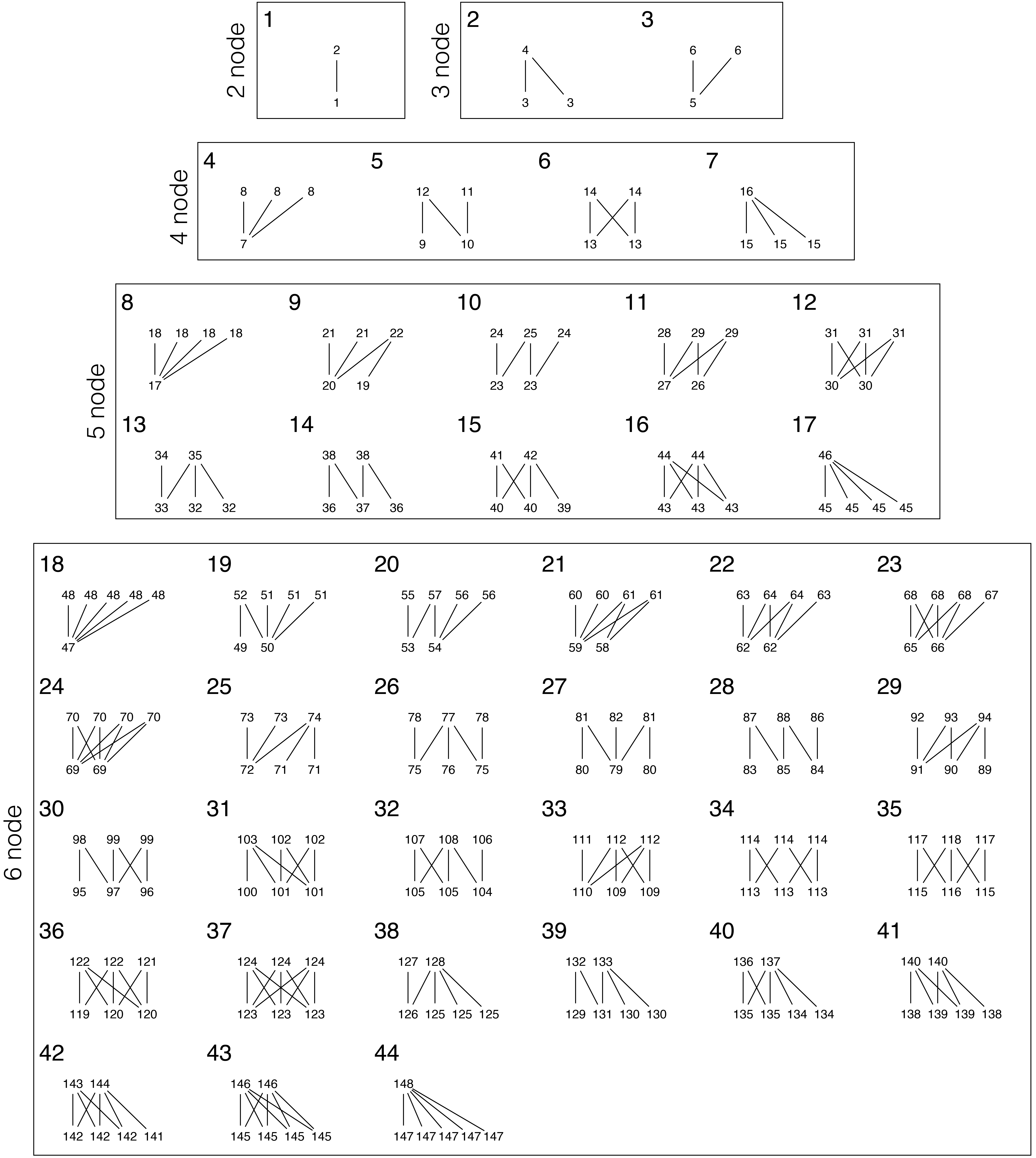
Here we introduce bmotif, a package for counting motifs, and species positions within motifs, in bipartite networks. Our code is primarily an R package, but we also provide MATLAB and Python code of the core functionality.
We describe the package and demonstrate how it can be used to conduct ecological analyses, using two examples of plant-pollinator networks. We first use motifs to examine the assembly and disassembly of an Arctic plant-pollinator community, and then use them to compare the roles of native and introduced plant species in an unrestored site in Mauritius.
bmotif will enable motif analyses of a wide range of bipartite ecological networks, enabling future research to characterise these complex networks without discarding important meso-scale detail on indirect interactions.